Pumping up the drops
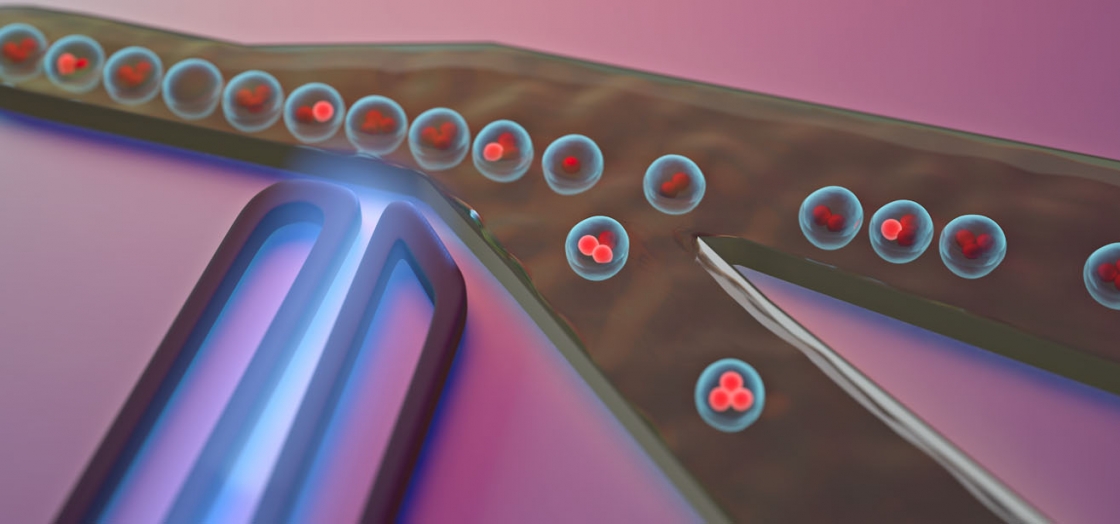
The group took a mathematical approach, Mukherjee said: if you increase the number of cells in each droplet, the probability that a droplet contains a fluorescing cell increases too.
“It's basically dumping out most of the non-fluorescing junk and selecting out the fluorescing population,” Mukherjee said.
Then, they repeat the flow cytometry process with the traditional single-cell per droplet approach—but this time, they sort out the material by more specific characteristics, such as a fluorescence lifetime or brightness.
The power of collaboration
To do that, they needed faster electronics and clean, precise tools. Those were all available in house at JILA, and the Jimenez Lab built their fast flow cytometry system completely at JILA.
JILA’s electronics shop was able to craft field-programmable gate array (FPGA) electronics which operate on a nanosecond scale—much faster than what they could order elsewhere. The clean room at JILA was used to fabricate all the microfluidic chips, so they were super clean. Being able to make everything in house also made this new system extremely cost-effective, Mukherjee added.
As a result, the Jimenez Lab enriched the proportion of fluorescing cells in their samples from 10% to 94%. They went from sorting 50 cells per second to about 2500 droplets per second—greater than a hundredfold improvement, Mukherjee said.
This type of system could make a difference not only to labs, but to anyone who needs to sort through a large library for a particular event, such as biomedical researchers who need to find the few abnormal cells in a pool of millions.
“We are trying to use it to approach fluorescent protein libraries but this is a very general approach to enrich any fluorescent event in a library of events.”
This study was published in Royal Society of Chemistry’s Lab on a Chip on February 21, 2020, and was supported by the NSF Physics Frontier Center Grant, and the NIH/CU Molecular Biophysics Training Program.
Written by Rebecca Jacobson