Protein molecular machines acting on DNA
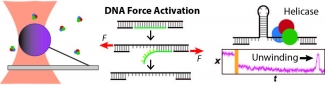
We apply optical traps and AFM to yield mechanistic insight into how enzymes move along and bind to DNA as well as the fundamental mechanical properties of DNA. We developed an actively stabilized, optical-trapping microscope featuring 1-bp resolution and applied it to the RecBCD helicase to show that RecBCD does not move in well-ordered discrete steps of either 1 or 4 base pairs (our anticipated result). We also developed a set of novel force-activated substrates that accelerated high-resolution studies of proteins that bind to and move along DNA, such as helicases. We used this technology to measure the on-rate for RecQ helicase binding to ssDNA, traditionally a difficult measurement using a single-molecule assay. We are currently integrating these advances to characterize the motion of helicases along DNA.
Measuring energy landscapes with a commercial AFM
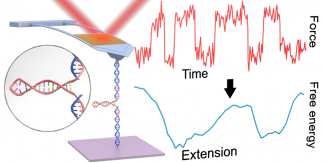
Reconstructing the full 1D free-energy landscape for folding of a biomolecule remains a pinnacle of SMFS measurements because the landscape is so information-rich compared to determining just a few landscape-describing parameters. However, such energy-landscape reconstructions have been achieved for only a few select nucleic-acid and protein structures, and only by a handful of leading labs using state-of-the-art optical traps. Integrating all of our technological improvements including modified cantilevers, we were able to characterize the dynamics of a short RNA hairpin from HIV on a commercial AFM and, in turn, to reconstruct its full 1D folding free-energy landscape.
Imaging protein-DNA interactions
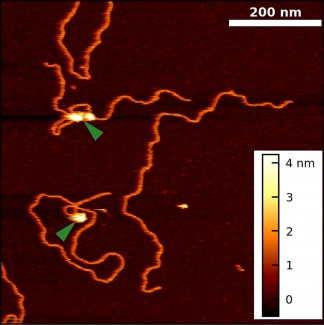
Historically, the biophysical insights derived from AFM imaging of DNA-protein complexes came with the caveat that the complexes are often dried and/or became partially trapped in three-dimensional configurations (“kinetically trapped”). These perturbations of the equilibrated structure on a substrate made it difficult to distinguish strand crossing from protein binding. To overcome these issues, we developed a simple 5-min protocol for depositing DNA and DNA-protein complexes onto mica in biologically relevant deposition conditions (10 mM MgCl2, 25 mM KCl). Importantly, our method yields images of DNA in liquid with its native physical properties [persistence length, rise per base pair, width, and helical pitch].
Using this protocol, we collaborated with Prof. Tom Cech (CU) to visualize Polycomb repressive complex 2 (PRC2) bound to DNA in a variety of protein-DNA complexes, including looped structures. These experiments yielded two unexpected results: PRC2 both compacts DNA and induces bends in DNA’s helical axis upon binding. Going forward, our improved technique can be extended to other protein-DNA systems and to more native-like PRC2 interactions (i.e., AFM imaging of nucleosome-PRC2 complexes).