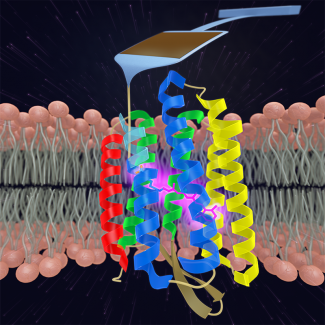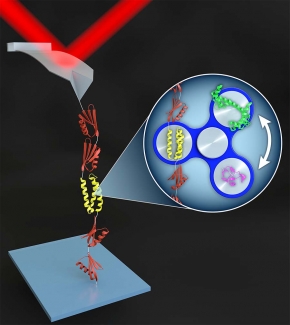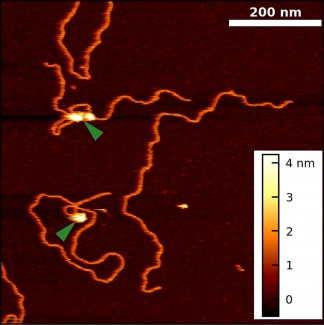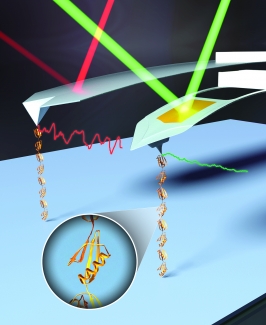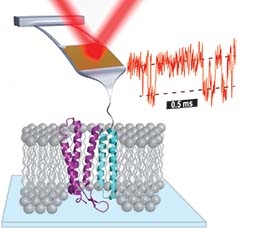
In the Perkins group, we develop and apply high-precision single-molecule techniques—atomic force microscopy (AFM) and optical traps—to address outstanding questions in a wide range of biological systems. Ongoing projects include elucidating the energetics and folding dynamics of membrane proteins embedded in their native lipid bilayer, characterizing the force needed to extract tubulin from a microtubule, probing the nanomechanics of myosin, and studying the motion of helicases along DNA. Our biological goals often motivate improvements in single-molecule techniques, such as developing focused-ion-beam-modified AFM cantilevers, an optical-trapping microscope with 1 Å stability in 3D, efficient site-specific conjugation strategies, and novel DNA- and protein-based constructs.