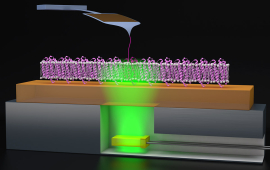
JILA biophysicists apply tools and concepts from physics to the understanding of living systems at the molecular level, particularly in molecular biophysics, single-molecule biophysics, and biological force standards.
The field of biophysics promises answers to important questions about the structure, dynamics, function, and interactions of biological molecules such as proteins and nucleic acids. JILA scientists are developing unique ways to precisely measure the structure and function of individual proteins and nucleic acids, both of which are important molecules for living organisms. JILA scientists are also pioneering new methods to image and measure real-time biochemical activities within living cells and tissues. These research endeavors advance our understanding fundamental biological processes and support biomedical scientists in developing new drugs and diagnostics.





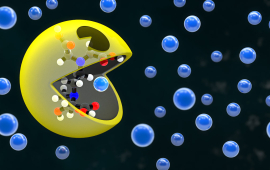
![The many different molecules trying to fill the binding site of octamethyl calix[4]pyrrole (omC4P)](/sites/default/files/styles/media_thumbnail_large/public/images/2025-04/weber_omc4p_thumbnail.jpg?h=fbf7a813&itok=e9ho_B28)